Perform group FDR estimation for different identification groups
FragPipe can be downloaded here. Follow the instructions on that same Releases page to launch the program. See here for help configuring FragPipe.
FragPipe can perform a group FDR estimation to calculate FDRs for each group separately. Then groups can be the number of enzymatic termini (i.e. 0, 1, 2), the protein evidence levels encoded in the FASTA file, and modifications.
Use the number of enzymatic termini or the protein evidence levels to group the PSMs
The group FDR approach supports most workflows. After loading a specific workflow, go to the MSFragger tab to select the group type.
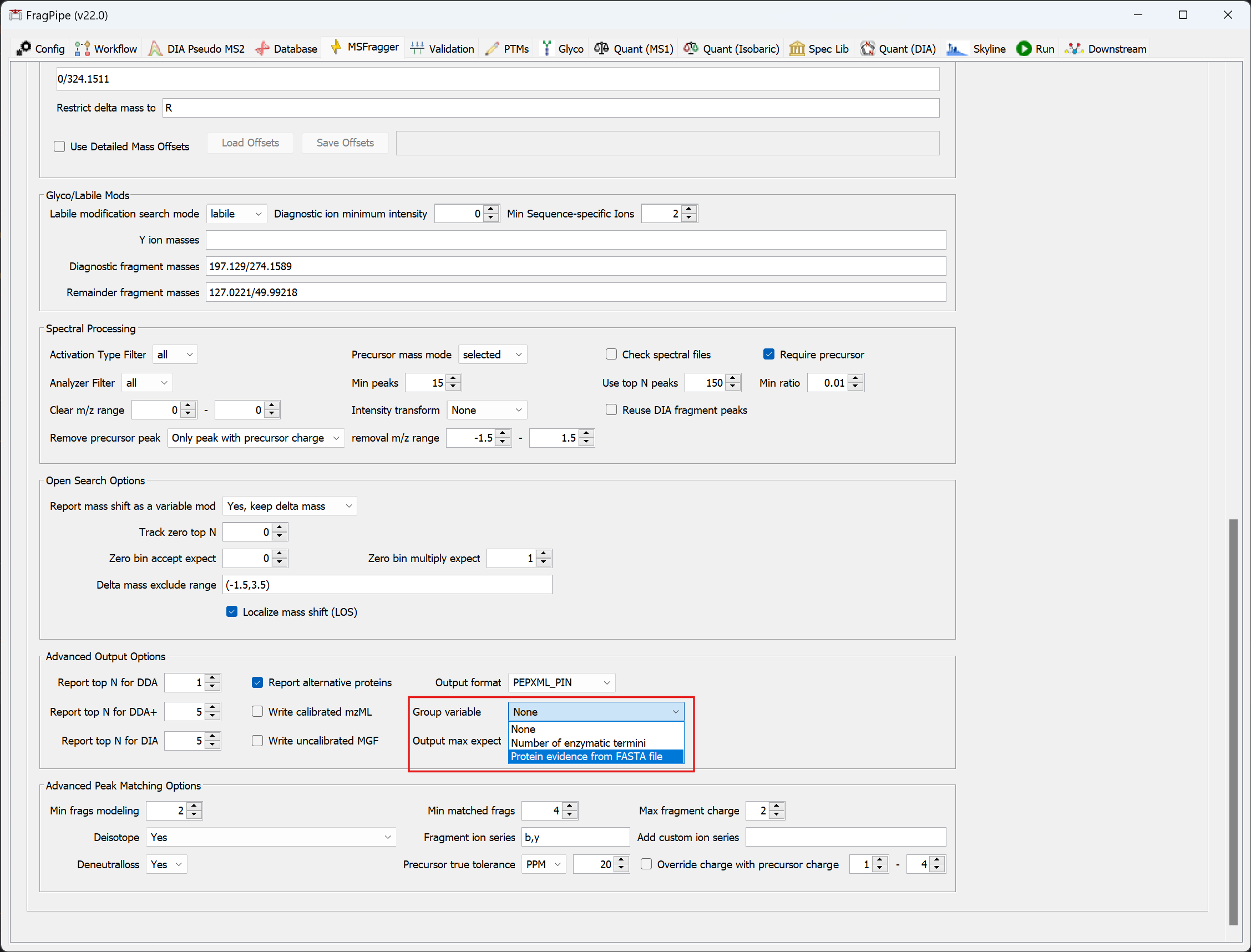
Then, go to the Validation tab, and add --group to the FDR Filter and Report command box.
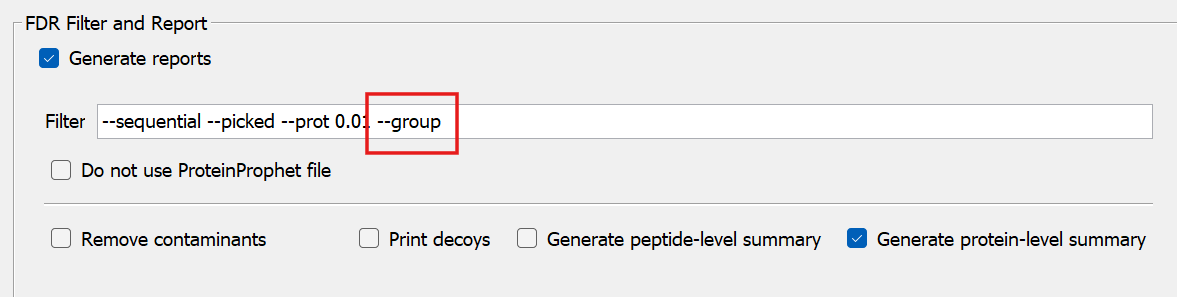
The rest of the part is the same as that in a normal FragPipe analysis.
To make FragPipe recognize the encoded protein evidence, the protein header in the FASTA file must have the PE= keyword. The numbers after PE= identify different groups. Here has a handy script to revise the protein headers.
Use the modifications to group the PSMs
In MSFragger tab, set the Group variable to None.
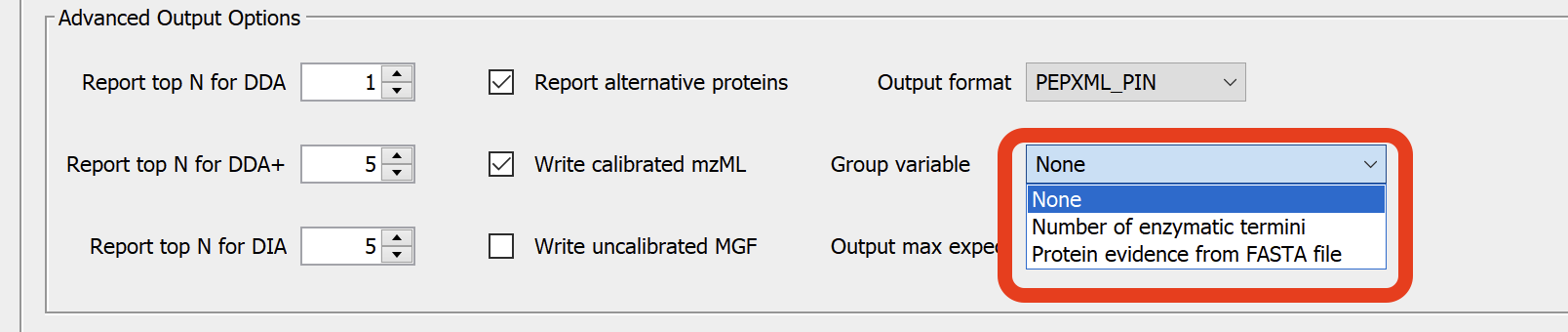
Go to the FDR Filter and Report panel in the Validation tab, add the modifications you want to group using the --mods flag. The following is an example.
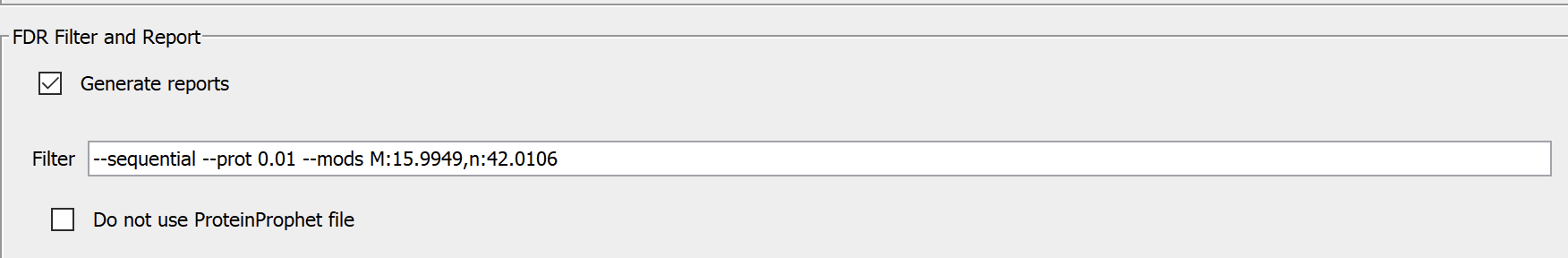
Philosopher will create 3 groups: unmodified peptides, peptides with the specified modifications only, and all other modifications. In the above example, the specified modifications are oxidation and protein N-term acetylation. Then, FDR will be computed separately for 3 groups: unmodified, specified common modifications, and rare modifications that are not specified.
Key Reference
Ferreira, H.J., Stevenson, B.J., Pak, H., Yu, F., Oliveira, J.A., Huber, F., Taillandier-Coindard, M., Michaux, J., Ricart-Altimiras, E., Kraemer, A.I., Kandalaft, L.E., Speiser, D.E., Nesvizhskii, A.I., Müller, M., Bassani-Sternberg, M. Immunopeptidomics-based identification of naturally presented non-canonical circRNA-derived peptides. Nature Communications 15:2357 (2024).